
CellBench: a new package helping researchers to select the best tool for interpreting single-cell datasets.
Much is still unknown about the trillions of cells in the human body. In the quest to better understand cells and the role they play in health and disease, a technique called single-cell sequencing has become a hot research field. Over the past five years there has been an explosion of new analysis tools for interpreting single-cell data. This has left researchers with hundreds of options to choose between in the challenging task of interpreting large and complex biological datasets.
With our close collaborators from WEHI, Luyi Tian and A/Prof Matt Ritchie, and in collaboration with the mixOmics team (Al Abadi), we have published in Nature Methods a first-of-its-kind data analysis platform to enable researchers to select the best tool for interpreting the overwhelming amounts of data generated by single-cell research. Importantly, we have designed our own gold standard single cell experiments to provide the community high quality data for computational methods development.
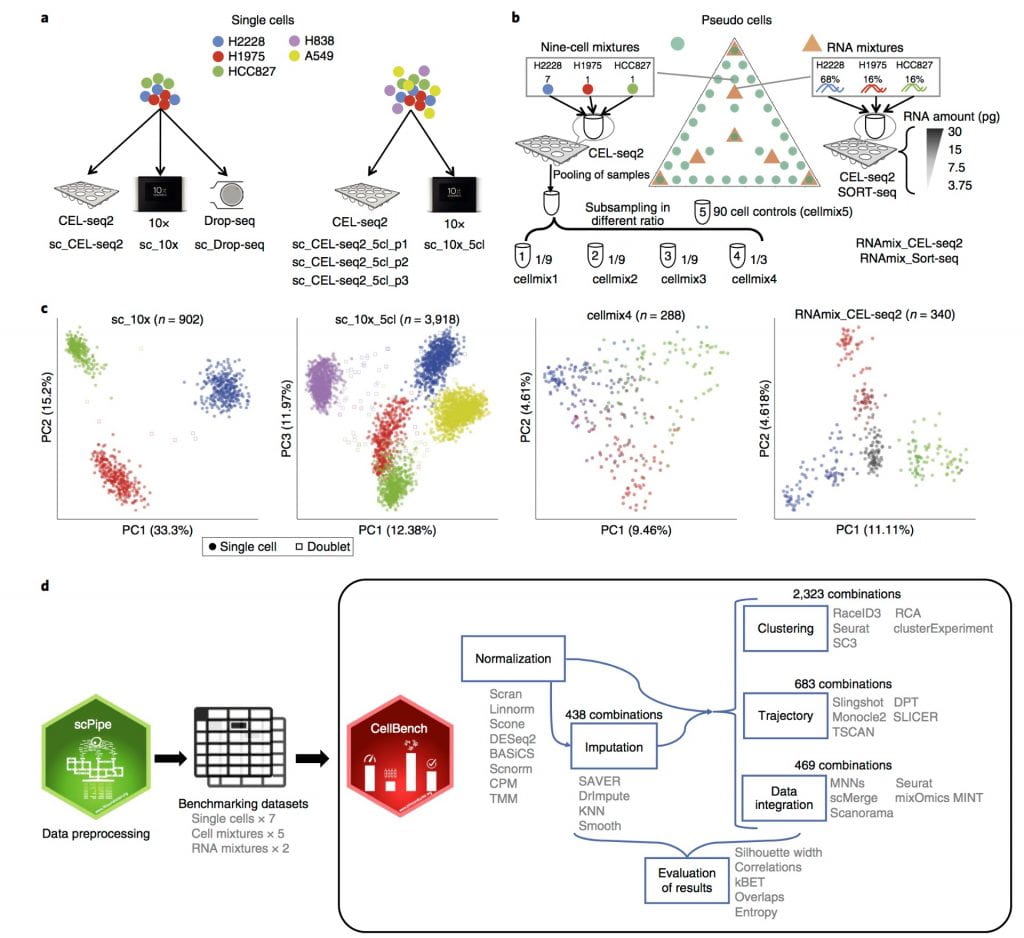
Choosing the right data analysis tool is crucial for avoiding misleading results or the incorrect biological interpretation of data. Cellbench fills this gap, by proposing a software and several gold standard datasets compares the performance of thousands of different single-cell analysis options, enabling researchers to identify the best method for the questions they wish to answer.
Development version of Cellbench is on Bioconductor at this link and on Shian (benchmarking single cell analysis methods) and Luyi’s (comparative analyses and generate the figures) gitHbub pages.
Reference:
The research was supported by the Chan Zuckerberg Initiative and the Australian Government’s National Health and Medical Research Council.
Categories